Addressing Difficult to Treat Bacterial Infections
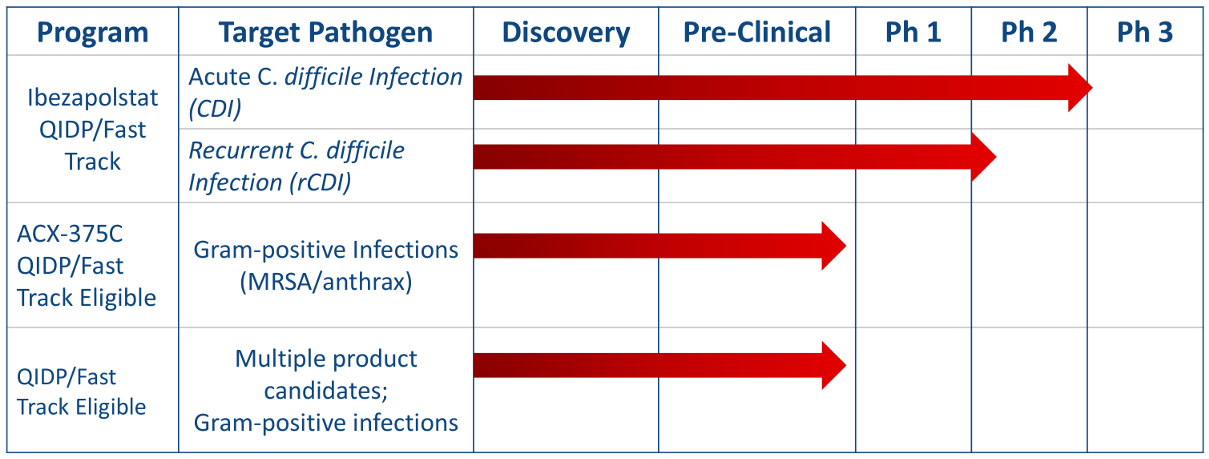
Acurx's approach is to develop antibiotic candidates that block an entirely new molecular target previously unexploited in medical research, DNA polymerase IIIC (Pol IIIC). Our R&D pipeline includes clinical stage and early stage antibiotic candidates that target Gram-positive bacteria for oral and/or parenteral treatment of infections caused by Clostridium difficile, Enterococcus (including vancomycin-resistant strains of VRE), Staphylococcus (including methicillin-resistant strains of MRSA), and Streptococcus (including antibiotic-resistant strains). Both of these antibiotic product candidates have demonstrated the ability to effectively block the DNA polymerase IIIC enzyme in early stage studies.
The CDC (Centers for Disease Control & Prevention) has designated C. difficile as a qualifying pathogen under the GAIN Act as an urgent threat highlighting the need for new antibiotics1 See “Qualified Infectious Disease Product (QIDP) Status by the FDA.” Further, the CDC has designated vancomycin-resistant enterococci as a serious threat level priority for new antibiotic development where, despite current treatment, a significant incidence remains and significant number of deaths occur annually.
A Scientific Roadmap for Antibiotic Discovery
A sustained and robust pipeline of new antibacterial drugs and therapies is critical to preserve public health
The Pew Charitable Trusts, May 2016 (accessed and excerpted July 2021); Deak D, Powers JH, Outterson K, Kesselheim AS, Progress in the Fight Against Multidrug Resistant Bacteria?: A Review of FDA Approved Antibiotics 2010-2015. Annals of Internal Med 31May2016
The State of the Field
Once a model of productivity and innovation, research efforts to discover new antibiotics have stalled. In the post-World War II period following Alexander Fleming’s breakthrough discovery of penicillin and the partnership between industry and government to produce this lifesaving drug on an industrial scale, new antibiotics were discovered and developed at a breathtaking pace. Such efforts led to dramatic advances in human health, as antibiotics were used to treat an increasingly wide range of infections while allowing for the evolution of the complex medical care that is now taken for granted, such as hip replacement, intensive care medicine, dialysis, and cancer treatment.
Drug discovery, the process of finding or designing molecules that could someday lead to new therapies, underpins drug development, the process of rigorously testing a therapeutic candidate for safety and efficacy in order to bring a new medication to market. The “golden age” of antibiotic discovery peaked in the 1950s, bringing forth lifesaving drug classes, such as erythromycin, vancomycin, and metronidazole. During this period, the pharmaceutical industry was the engine of innovation as nearly every major company maintained an active research and development (R&D) program in antibiotic research.3 Yet, after the initial rush of new compounds—many isolated from actinomycetes, bacteria that are mainly found in soil—new starting points for antibacterial drugs became harder to find, and the scientific hurdles more apparent.
New discoveries dropped precipitously from the 1980s onward. As a result, the development of antibiotics has declined, with new Food and Drug Administration (FDA) approvals for these drugs falling from 29 during the 1980s to nine in the first decade of the 2000s. All antibiotics approved for use in patients today are derived from a limited number of types, or classes, of antibiotics that were discovered by the mid-1980s. This is even more concerning than the decline of drug approvals because resistance to one antibiotic often leads to resistance to multiple antibiotics within the same class. While drugs can be categorized or classified in a variety of ways, for the purposes of this document, antibiotic classes are based on similarities in chemical structure.

Why Do We Need New Classes of Antibiotics?
Antibiotics can be categorized based on similarities in their chemical structures (i.e., antibiotic classes).
Resistance to one antibiotic often leads to resistance to multiple antibiotics within the same class.
Source: Adapted from Lynn L. Silver, “Challenges of Antibacterial Discovery,” Clinical Microbiology Review (2011)
© 2016 The Pew Charitable Trusts
In the face of this mounting crisis, efforts to revive and improve the likelihood of successful drug discovery are essential. The U.S. government, industry, and the public health and medical communities all agree that new regulatory policies and economic incentives are critically needed to rebuild a robust pipeline of antibiotics.
References on file: Acurx Pharmaceuticals; July 2021